Functional and Systems Biology
Visualizing the Biological Dynamics of Cells
EMSL’s live cell imaging platforms provide real time, 3-D microscopy
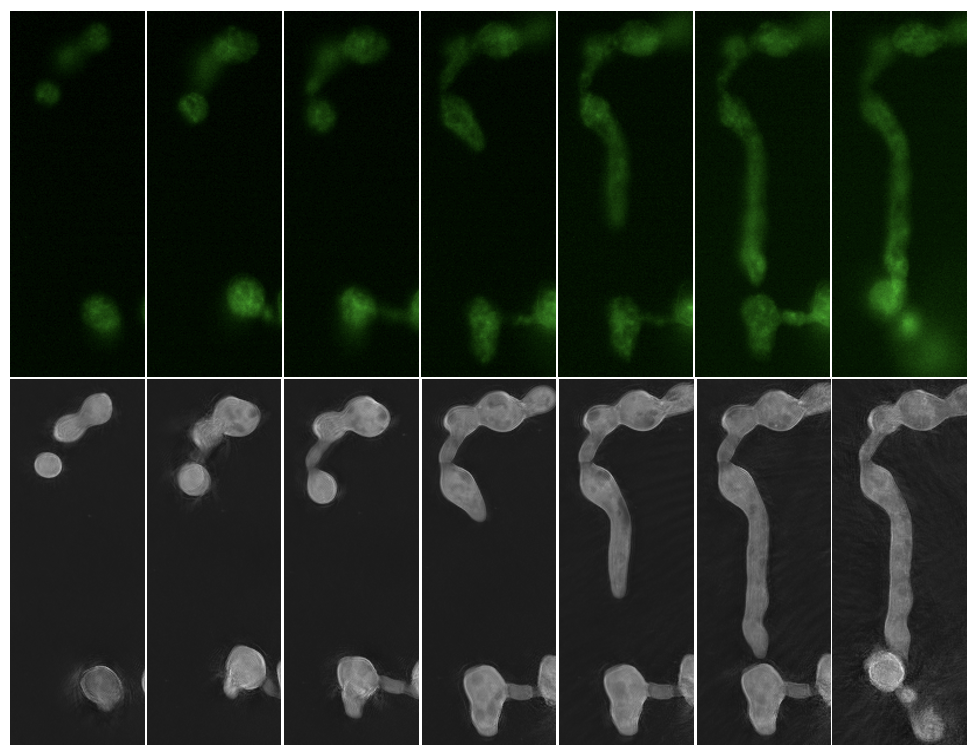
Live cell microscopy platforms like NanoLive allow researchers to take a deeper look inside single cells and whole cell populations. These resources are available to the scientific community through EMSL’s open calls for proposals. (Microscopy image provided by Erin Bredeweg)
Register to attend a free webinar at noon PST on Wednesday, Jan. 24 to learn how access and apply EMSL's 3-D platforms for live microscopy. These resources are available through EMSL's open calls for proposals.
Live cell imaging has emerged as a foundational component to biological research.
Using a variety of methods of time-lapse, 3-D microscopy, live cell imaging allows researchers to visualize and follow the dynamic processes of single cells, whole cell populations, and subcellular activity.
Unlike static experiments where cells are chemically fixed into a behavioral or structural state at a specific time, live cell imaging platforms provide insight into spatiotemporal cellular changes in real time. As a result, these powerful methods enable a more comprehensive, physiologically realistic view of cellular processes.
At EMSL, the Environmental Molecular Sciences Laboratory, a Department of Energy Office of Science user facility, several live cell imaging platforms are available to the research community through open calls for proposals. If awarded, principal investigators and their research teams gain access to requested scientific resources and EMSL staff expertise at no cost.
These platforms include the NanoLive microscopy platform, which is designed for repeat observations over time on bacterial and eukaryotic cells, and the Lattice Light Sheet microscope, which enables the imaging of whole small organisms down to single molecules.
NanoLive
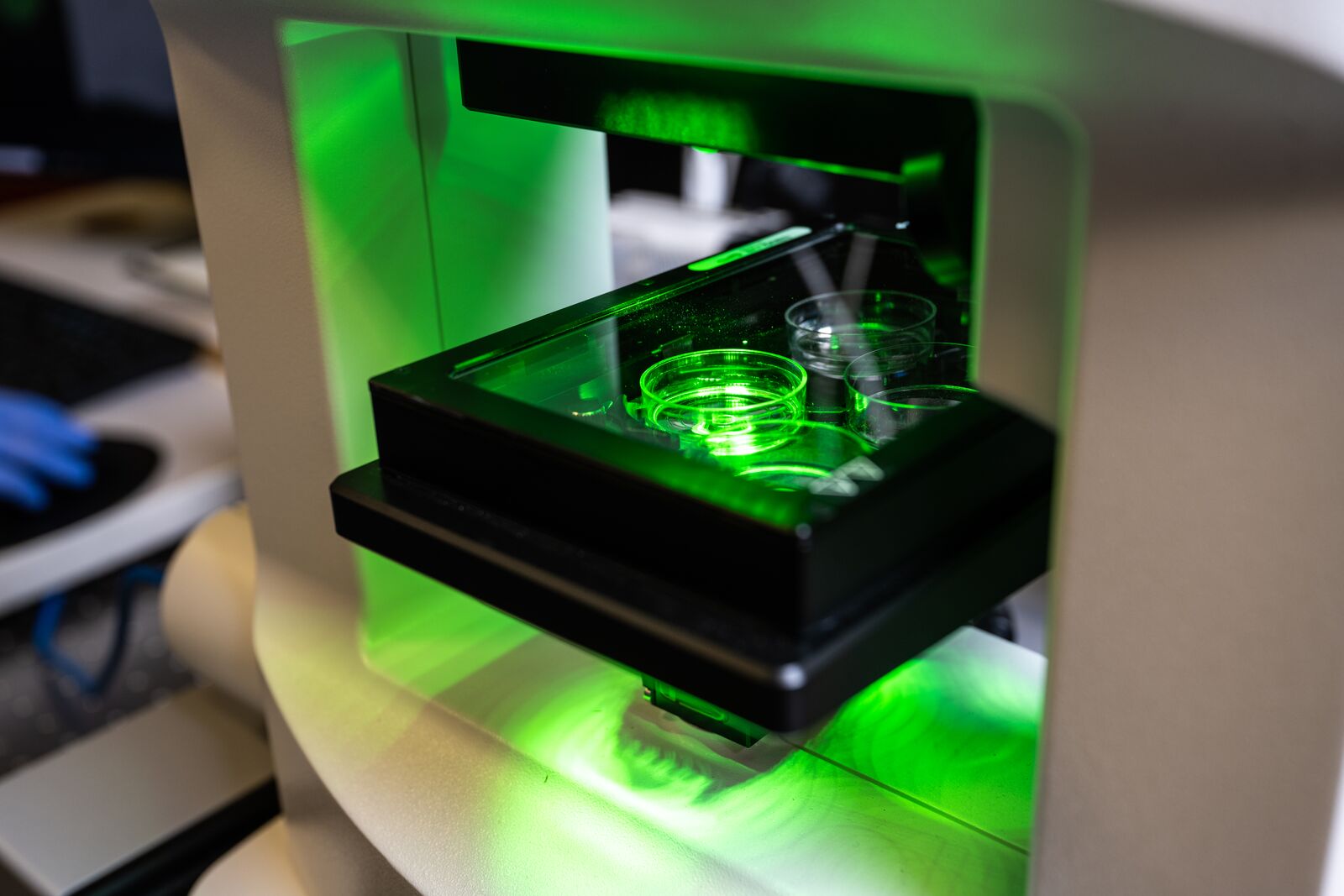
The NanoLive platform is a noninvasive and label-free tool for long-term observations of living cells. This automated instrument creates 3-D images of samples with the use of holographic tomography and fluorescent signal images.
As a newly offered EMSL capability, researchers can apply NanoLive to study protein function and the localization of cells to observe bacterial, plant, or fungal cells in real-time and to examine complex cellular phenotypes. By using NanoLive, researchers can capture multiframe, multilayer images of cell–cell interactions, phenotypic responses to growth conditions or chemical signals, and organellar dynamics.
“Researchers who are awarded instrument time at EMSL can use NanoLive’s rapid data capture and low phototoxicity for timelapse observations, which are challenging on standard microscopy platforms,” said Erin Bredeweg, an EMSL biologist who is the primary contact for NanoLive. “With the ability to capture multiple, parallel conditions, the NanoLive platform is ideal for extended observation, all with room for biological replication.”
NanoLive’s 3-D holography provides a unique window into living cells, while the added layer of fluorescence data collection makes it compatible with EMSL’s activity-based probes, fluorescently tagged proteins, and stains.
For data collection from live cells, users send live cultures to EMSL, as well as any experimental specifications for media, temperature, or timing of observations. EMSL scientists communicate with the research team about preculturing conditions, and the best setting for the experiment, including imaging replicates (up to 60 wells). Endpoint data can be used for 3-D holographic construction and timelapse observations.

Lattice Light Sheet
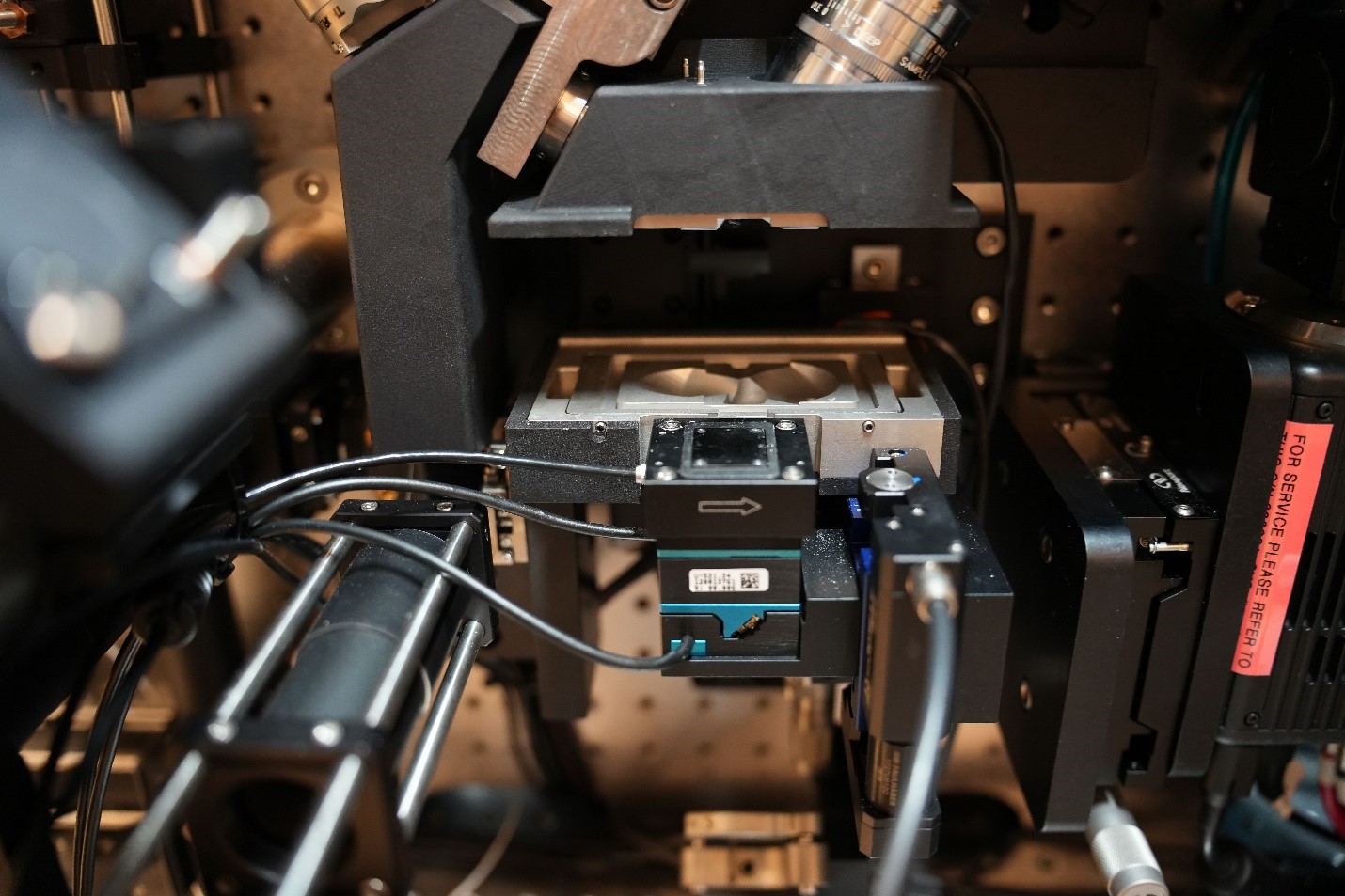
The Lattice Light Sheet microscope uses a modified version of light sheet fluorescence microscopy that combines a high spatiotemporal resolution, imaging speed, and sensitivity to capture 3-D images of organisms down to single molecules.
Nobel Laureate Eric Betzig developed Lattice Light Sheet microscopy in the early 2010s to be a “high-speed, high-resolution, low-impact tool that can look deep inside biological systems,” noted EMSL chemist Dehong Hu, who is the instrument contact for the microscope.
With the high speed of the Lattice Light Sheet microscope, samples are less likely to be exposed to phototoxicity, which has been a common challenge with fluorescence microscopy. With its fast image capturing capability, the Lattice Light Sheet microscope can also be used for hours or days, unlike the restrictiveness of traditional fluorescent microscopy instrumentation.
During experiments, Lattice Light Sheet microscopy uses thin sheets of light to create an optical lattice and to generate a series of images. The images can provide information about dynamic biological processes, including key protein and subcellular structures in plant cell tissues or microbial communities.
At EMSL, researchers who have been awarded access to use the Lattice Light Sheet microscope can connect with Hu before creating samples so that he can provide guidance on sample preparation. Hu has more than 20 years of experience in single-molecule fluorescence imaging and works with users on proposals related to fluorescence in situ hybridization, single-molecule imaging, and single-cell analysis.
Hu said that sample preparation is very different from other microscopes. The main difference is that the sample is immersed in an aqueous solution. With regular microscopes, samples are usually on microscope slides or in glass bottom dishes.

Researchers interested in using the Lattice Light Sheet microscope or NanoLive are encouraged to reach out to Alex Beliaev, the leader of EMSL’s Cell Signaling and Communications Integrated Research Platform (IRP) before submitting a letter of intent (LOI) to a proposal call. By engaging with IRP leaders before submitting an LOI, principal investigators and teams can address research questions and identify the instrumentation that is best suited for the project.

