Revealing the Stress Secrets of Plants One Cell at a Time
Researchers used a new approach to measure thousands of proteins from single plant cells experiencing water deficit stress, revealing new insights into cell-type specific impacts and driving innovations

A team of researchers applied a highly sensitive workflow that allowed them to measure more than 3,000 proteins within single cells (protoplasts) of Arabidopsis thaliana leaves under normal and reduced watering conditions. (Photo courtesy of dra_schwartz, iStock)
The Science
Water deficits add stress to plants, which in turn alters how they grow. In studying what occurs at the cellular level when plants experience water scarcity, most studies measure many cells at once and miss differences in individual cells. In this study, a team of researchers applied a highly sensitive workflow that scaled sample processing to tiny volumes, allowing them to measure more than 3,000 proteins within single cells (protoplasts) of Arabidopsis thaliana leaves under normal and reduced watering conditions. The protein "fingerprints" of individual cells were found to shift with water deficit stress, and proteins could be grouped into distinct clusters that aligned with their shared functions.
The Impact
Knowing how single plant cells respond to water deficit stress can help scientists connect stress to specific biological functions and, over time, guide efforts to improve crop resilience—including crops grown for bioproduct development. This study advances science because the authors identify and measure the proteins (the cell's working molecules) from a single cell, one cell at a time, rather than averaging across tissues composed of many types and quantities of cells. It is the first application of a workflow approach referred to as "label-free proteomics" at the single-cell level to plant systems. This approach reveals the underlying cell-to-cell variation and implicates the ribosome (the cell's protein factory) as a major player in water deficit stress. This new approach could be used in future larger-scale experiments to track which cell types or cell states drive stress tolerance and to test how genetic modifications or treatments shift those responses.
Summary
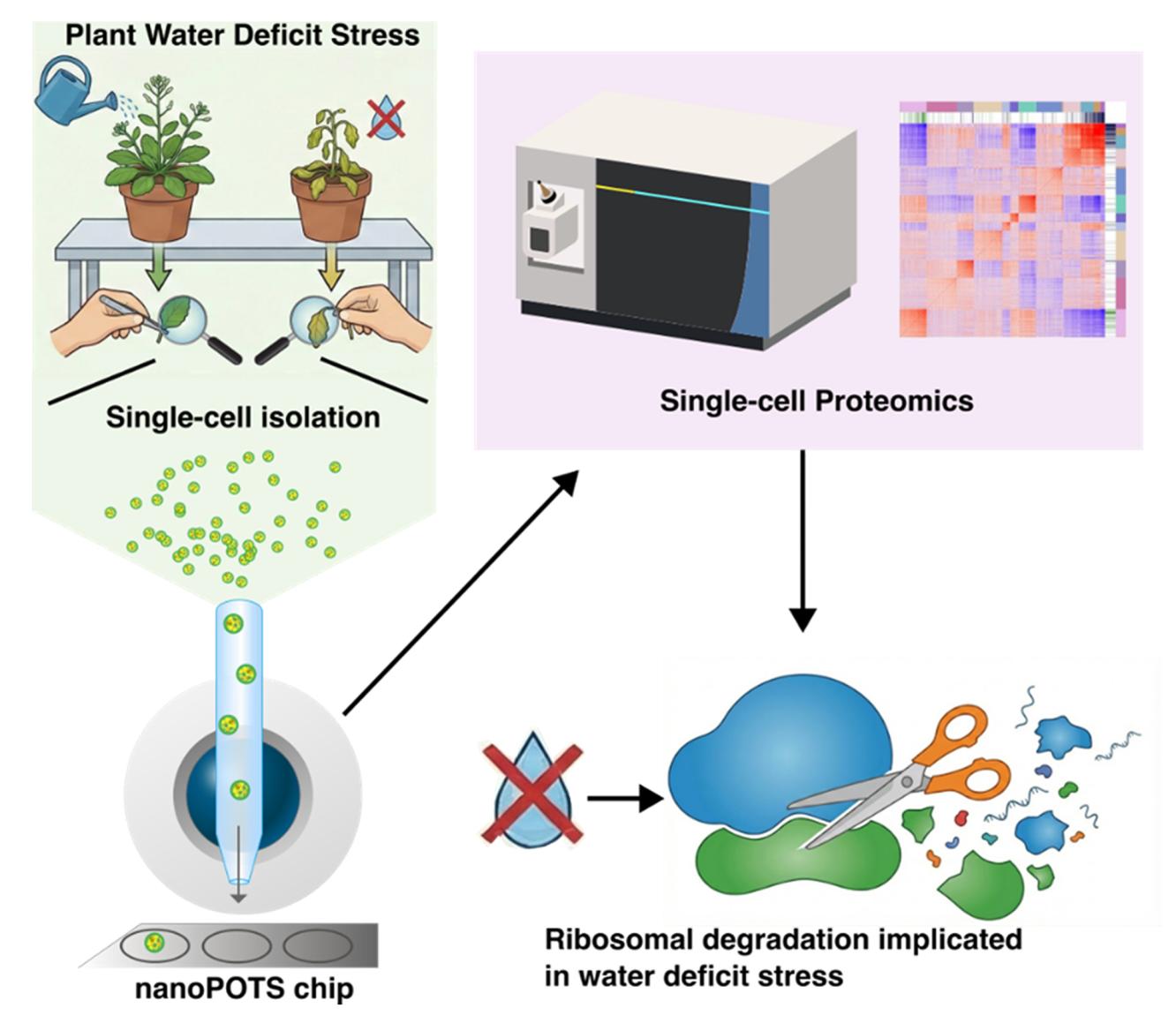
Plants are made of many kinds of cells, and even cells of the same type can respond differently to stress. Most "single-cell" plant studies have focused on ribonucleic acid (RNA) sequencing, which provides a picture of what a cell might do, not necessarily what it is currently doing or how it is responding to an external stimulus. This study demonstrates how thousands of proteins (the molecules that carry out most cell functions) can be measured in single cells from Arabidopsis thaliana leaves. Scientists at the Environmental Molecular Sciences Laboratory (EMSL), a Department of Energy Office of Science user facility, first separated cells from plant leaves using a technique called "protoplasting," and then they sorted them using a cellenONE sorter. The team prepared the cells for analysis using EMSL's Nanodroplet Processing in One Pot for Trace Samples (nanoPOTS) platform, and then they applied mass spectrometry techniques to measure the proteins that were within the cells. Using this workflow, more than 3,000 proteins could be measured with high reproducibility from a single cell. This approach was applied to more than 100 single cells from plants grown with normal watering or under water deficit conditions. Cell-type specific changes in protein abundances could be detected, and protein "co-variation" analysis (which identifies proteins that rise and fall together across cells) revealed not only new protein functions but how protein relationships could shift during water deficit stress.
Contacts
James M. Fulcher
Pacific Northwest National Laboratory,
EMSL
james.fulcher@pnnl.gov
Ljiljana Paša-Tolić
Pacific Northwest National Laboratory,
EMSL
Ljiljana.PasaTolic@pnnl.gov
Amir H. Ahkami
Pacific Northwest National Laboratory,
EMSL
amir.ahkami@pnnl.gov
Funding
This work was supported by the Department of Energy (DOE) Office of Science, Biological and Environmental Research program's Biopreparedness Research Virtual Environment project, a Laboratory Directed Research and Development award from Pacific Northwest National Laboratory, and a project award from the Environmental Molecular Sciences Laboratory, a DOE Office of Science User Facility sponsored by the Biological and Environmental Research program.
Publication
J.M. Fulcher, et al. "Single-cell proteomics of Arabidopsis leaf mesophyll reveals dynamic protein responses to water-deficit stress." Genome Biology (2026). [DOI: 10.1186/s13059-025-03919-6]